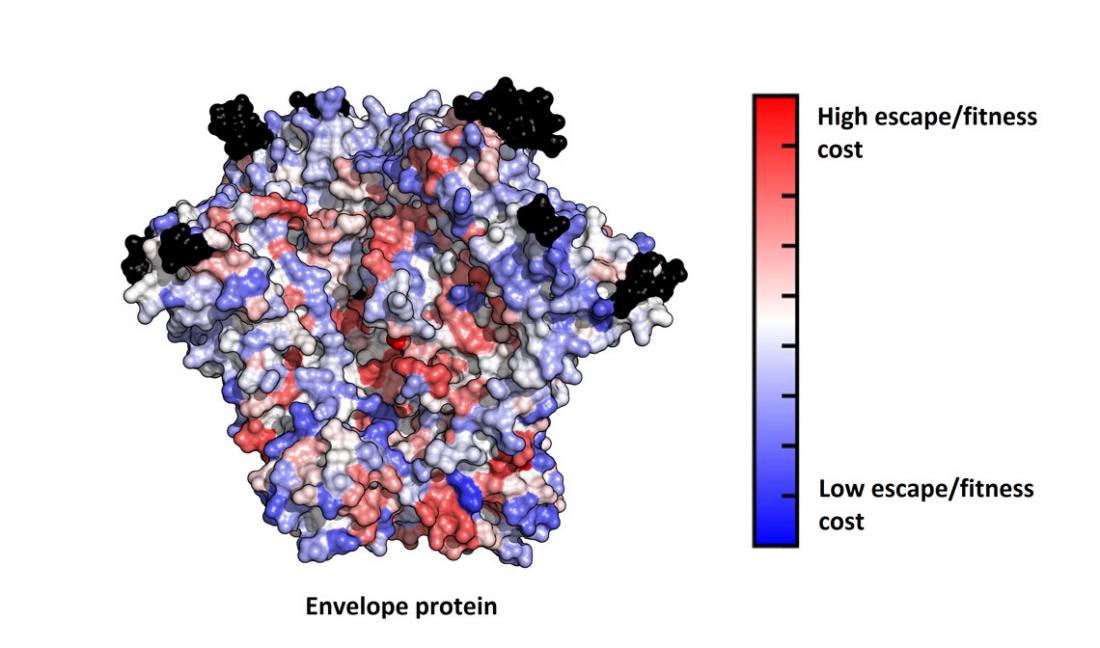
Potential weak spots on the surface of the HIV envelope protein (red) represent targets for rational vaccine design. Mutations at these positions are anticipated to be deleterious for viral fitness, whereas those at the blue regions are expected to be more benign. Hence, it may be effective to design a vaccine that forces the virus to mutate at the red positions.
Researchers in Hong Kong and the United States have identified potential weak spots in the part of the human immunodeficiency virus (HIV) genome that codes for proteins that help the virus attach to human cells. The team used a machine learning approach to make the finding, which could help guide vaccine development.
More than 31 million people are living with HIV worldwide. While antiretroviral drugs can help suppress the virus, an effective vaccine or cure does not exist. HIV vaccine development is challenging because the virus mutates rapidly and mutations in HIV surface proteins, called envelope proteins, can make them invisible to the body’s immune system.
“It is a very complex problem and a core reason why these viruses are so difficult to combat,” explains computational biologist Matthew McKay of Hong Kong University of Science and Technology (HKUST).
To address these challenges, McKay and his team, in partnership with colleagues at the Massachusetts Institute of Technology in the US, applied big data machine learning methods to publicly available amino acid sequence data to search for weak spots in certain regions of the HIV genome.
Their method involves mapping the so-called ‘fitness landscape’ of a virus: the relationship between the virus’s genomic sequences and its ability to assemble, replicate and propagate infection. By discovering which genomic regions are most critical for reproduction, researchers can design vaccines to elicit antibodies that drive gene mutations in those areas, inhibiting the virus’s ability to spread.
Knowledge of the genomic region that codes for HIV envelope proteins is a key to vaccine design, since they are the core targets of antibodies. Due to their high diversity, the fitness landscape of these proteins had remained unknown. Using machine learning, the researchers were able to map the fitness costs of mutations in specific parts of the genomic sequence encoding the HIV envelope polyprotein called gp160, revealing potential vulnerable regions in this protein.
“This knowledge helps us understand which mutations appear most likely to allow the virus to escape from vaccine-induced antibodies, which can possibly assist in the design of new vaccines with enhanced efficacy,” says Raymond Louie, a member of McKay’s HKUST team and a lead author of the study published in the Proceedings of the National Academy of Sciences.
Fitness landscape modelling can be applied to vaccine and drug design for other viruses and pathogens such as hepatitis C, which infects around three million people each year.
For further information, contact:
Professor Matthew McKay
Depts. of Electronic and Computer Eng, and Chemical and Biological Eng.
Hong Kong University of Science and Technology
E-mail: [email protected]
Dr Raymond Louie
Kirby Institute
University of New South Wales
E-mail: [email protected]