“Clock genes” turn on and off, or “express”, in rhythmic patterns throughout the body to regulate physiological conditions and behaviour. When and how these genes express, especially in tissues outside the brain, is still poorly understood. Until now, scientists have lacked sufficient means to simultaneously monitor gene rhythms in specific tissues in freely moving subjects.
Bioluminescence is a technique that often involves modifying certain genes. When the target gene is switched on, it also expresses an inserted gene, leading to the emission of a light signal. Every time the target gene turns on, light is emitted. This technique has been successfully used to monitor gene expression in fully anaesthetized mice. However, anaesthesia is believed to affect the expression of clock genes. Other techniques not requiring anaesthetization have their own limitations.
A team of scientists from Hokkaido University in Japan developed a new imaging technique that allowed them to monitor the expression of clock genes, including the per1 gene, in multiple tissues in moving fully conscious mice. When per1, which is known to be important for maintaining circadian rhythms, turns on in the transgenic mice, a light signal is emitted.
-------------------------------------------------------------------------------------------------
(Image left) In vivo live imaging system in freely moving mouse.
The system consists of two newly developed programs that automatically identify the 3D positions of target areas based on the signals from the scintillators (shown as white dots), and determine the intensity of Per1-luc bioluminescence in those target areas (shown as green-yellow circles).
--------------------------------------------------------------------------------------------------
To track the movement of the mouse, scintillators that fluoresce were attached to the back and head of the mice. The team developed a software program that was able to detect the three-dimensional position of the scintillators in the freely moving mice based on images received from two separate cameras placed in the mouse cage. The team also developed a set of algorithms that allowed them to identify the intensity of the bioluminescent signals from target tissues (olfactory bulb, right and left ears and cortex, skin) despite the movement of the mice.
“Using the present system, we observed robust circadian rhythmicity in per1 expression at six different areas in the bodies of freely moving mice,” the researchers report in their study published in the journal Nature Communications. Per1 expression was at its peak in all six areas at the onset of mouse daily activity. When the Hokkaido team artificially shifted the hours of night and day for the mice, such as might happen when shifting time zones, per1’s rhythmic expression became desynchronized for one day in the different areas and then synchronized again.
While further improvements are still required, the new monitoring technique could be widely applied to many areas of biomedical research, as well as in applications beyond medicine, the researchers say.
This work was supported by Creation of Innovation Centers for Advanced Interdisciplinary Research Areas Program, Ministry of Education, Culture, Sports, Science and Technology, Japan.
Contacts:
Associate Professor Toshiyuki HAMADA
Graduate School of Medicine
Hokkaido University
[email protected]
Mr. Naoki NAMBA (Media Officer)
Global Relations Office
Office of International Affairs
Hokkaido University
Email: [email protected]
Tel: +81-11-706-8034
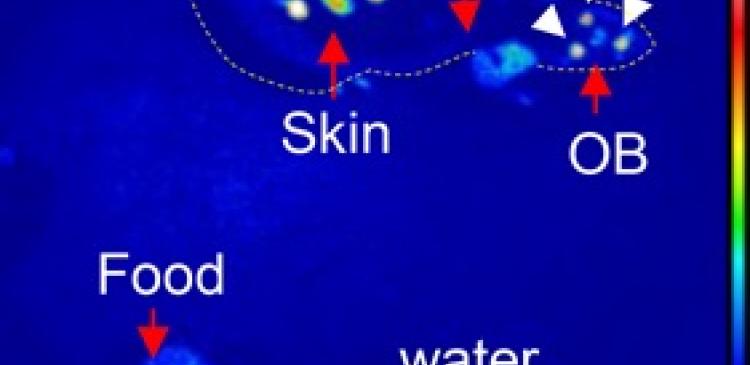
(Image below)
Representative images of Per1-luc mice.
The fluorescence signal from the scintillators (white arrow heads) and bioluminescent signals from the target areas (olfactory bulb, skin) are acquired by cameras. The dotted line shows the shape of the mouse body. (Hamada et al., Nat Commun. 2016 Jun 10)