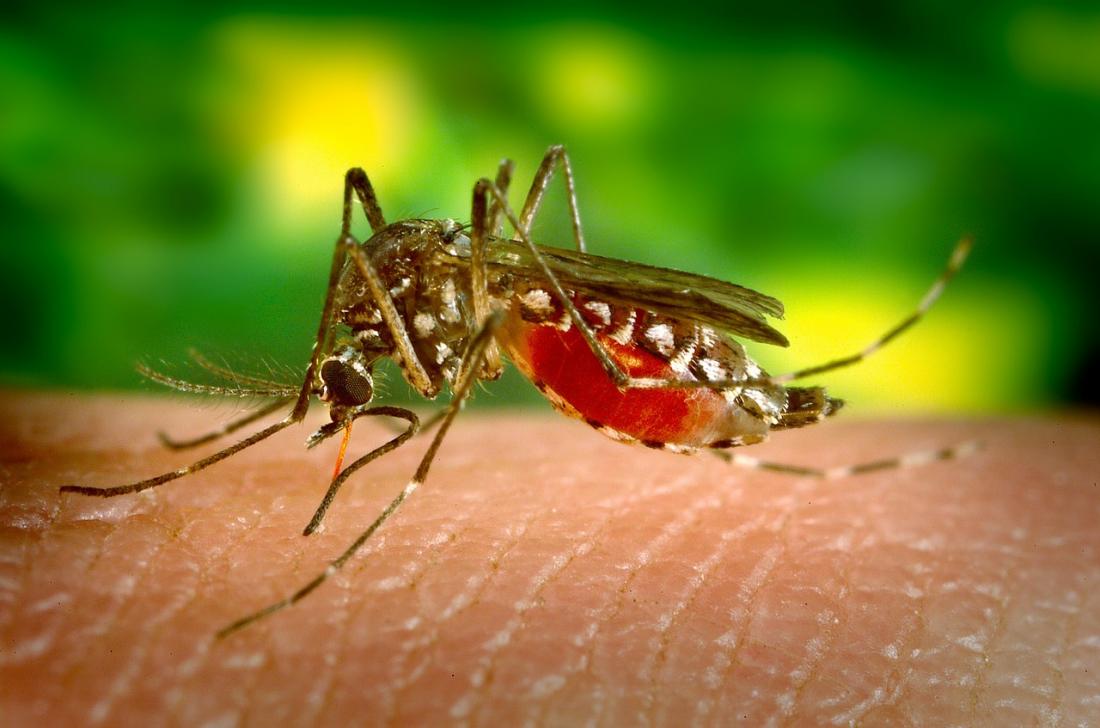
The results highlight the difficulties in predicting dengue epidemics but also emphasize the need to reconstruct multiple models to more fully represent all potential epidemic trajectories, as when predicting climate changes. [Image provided by Asia Research News]
An international team has been improving dengue infection outbreak predictions, highlighting the challenges researchers face when attempting to build successful forecasting models.
Dengue is a mosquito-borne viral infection most commonly found in the Caribbean, the Americas, and South East Asia. The infection is usually mild, but can also cause severe flu-like symptoms and can be life-threatening. According to the World Health Organization, up to 100 million infections are estimated to occur annually in over 100 countries, putting almost half of the world’s population at risk.
Major dengue epidemics occur every two to five years depending on climate fluctuations. There is currently no definitive treatment, so prevention is tantamount to reducing the numbers of people affected. If scientists were able to predict an epidemic, healthcare providers would be better positioned to prevent and control it.
The US Pandemic Prediction and Forecasting Science and Technology Working Group established the Dengue Forecasting Project in 2015. Sixteen teams from around the world were given the same dengue incidence and climate data for Iquitos, Peru and San Juan, Puerto Rico in order to understand the universal drivers of Dengue epidemics. The researchers combined disease and climate data from previous years and the current season into complex models and tried to predict how the season would progress prior to the occurrence of outbreaks.
“Developing dengue forecasting models has so far been challenging,” explains Hokkaido University information scientist Matteo Convertino, who led one of the sixteen teams in the project. “Forecast targets often vary erratically, making it difficult to compare and evaluate them, and environmental variables are often tied weakly to population health targets.”
Matteo Convertino at the podium of WHO/PAHO in Washington, D.C.
The Dengue Forecasting Project attempted to overcome these challenges by providing three clear target predictions: maximum weekly number of cases, the week when maximum cases occur, and the total number of cases in a season.
The models were then tested using another known data set from the same locations, to see how accurately they predicted the epidemic targets. None did very well, but performances generally improved as the infection season progressed. Models that produced an ensemble of forecasts by running through the data several times with slightly different starting points robustly outperformed models that conducted only individual forecasts. Some of the models were skillful in producing forecasts for individual seasons as far as up to four months in advance.
The results, published in the journal PNAS, highlight the difficulties in predicting dengue epidemics, even in endemic areas with clear seasonal transmission patterns. However, the results also emphasize the need to reconstruct multiple models to more fully represent all potential epidemic trajectories, as when predicting climate changes. This is also useful for exploring competing hypotheses of how epidemics work.
“The challenges are formidable, but through forecasting projects like this one, the community can move this research forward, translating the research into public health tools that can transform the way we prepare for and respond to epidemics,” the researchers conclude.
Funding:
Several departments in the U.S. Federal Government (Department of Health and Human Services, Department of Defense, Department of Commerce, and the Department of Homeland Security) are acknowledged to have joined together, with the support of the Pandemic Prediction and Forecasting Science and Technology Interagency Working Group under the White House National Science and Technology Council, to design the infectious disease forecasting project with the aim of galvanizing efforts to predict epidemics of dengue.
Contacts:
Associate Professor Matteo Convertino
Nexus Group
Graduate School of Information Science and Technology
Global Station for Big Data and Cybersecurity, GI-CoRE
matteo[at]ist.hokudai.ac.jp
Naoki Namba (Media Officer)
Institute for International Collaboration
Hokkaido University
Tel: +81-11-706-2185
Email: en-press[at]general.hokudai.ac.jp