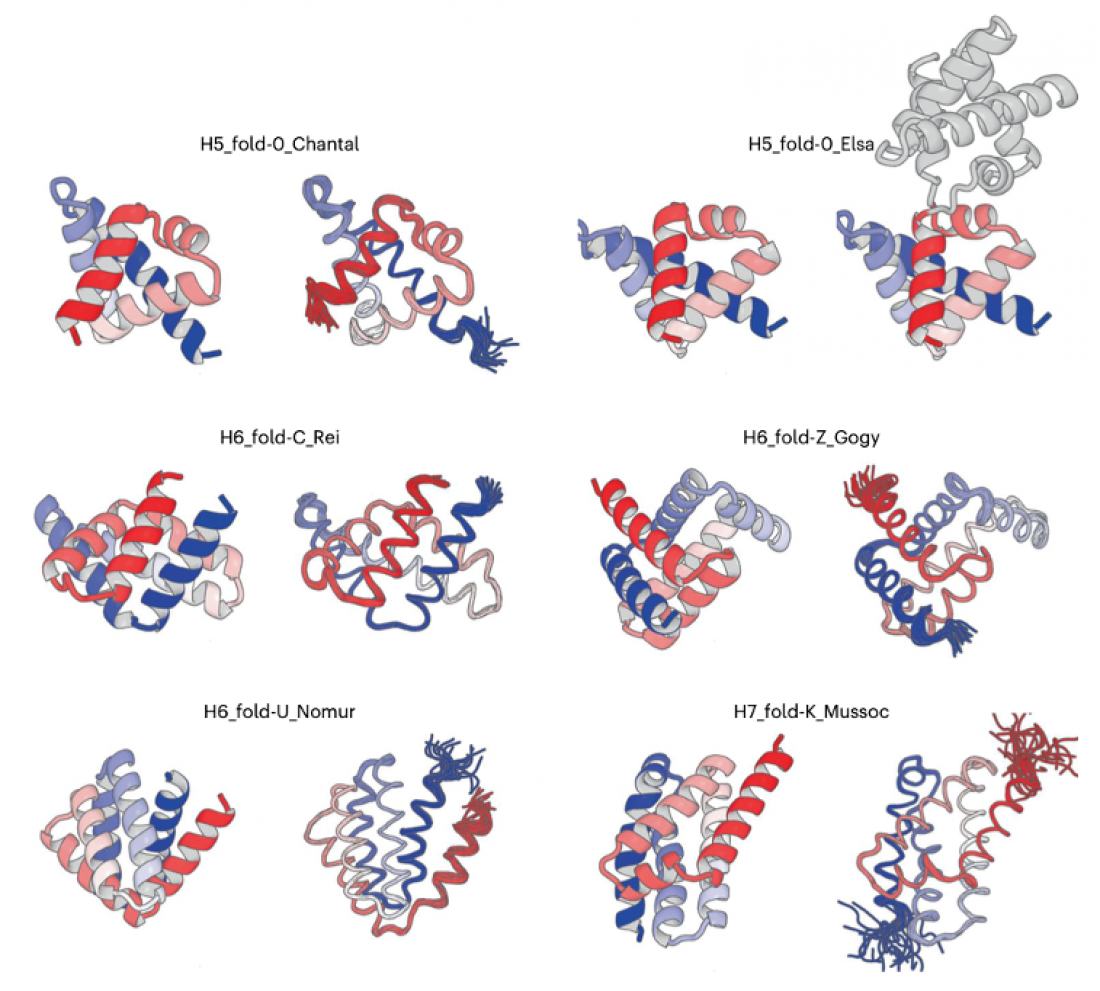
De novo designed all-a proteins with complicated shapes: H5_fold_0_Chantal, H5_fold-0_Elsa, H6_fold-C_Rei, H6_fold-Z_Gogy, H6_fold-U_Nomur, and H7_fold-K_Mussoc are presented. For each de novo designed protein, the computational model is shown on the left, and the solved experimental structure is on the right.
Proteins fold into unique three-dimensional structures based on their amino acid sequences, which then dictate their function. Although significant progress has been made in de novo protein design, the ability to design complicated all-α proteins, where α-helices are non-parallelly arranged within the three-dimensional structures, was lacking. “Artificially designed proteins mostly show simple structures, but Nature presents us with complicated ‘designs’” said Prof. Nobuyasu Koga of Exploratory Research Center on Life and Living Systems (ExCELLS) at National Institutes of Natural Sciences (NINS). This gap drove the team to seek a method to design such complicated all-α proteins.
The team began by examining structures deposited in the Protein Data Bank (PDB) and identified 18 typical helix-loop-helix motifs. They then demonstrated that a broad spectrum of all-α protein tertiary structures, ranging from simple to complicated, can be computationally generated by combining these identified typical motifs and canonical α-helices. “It is surprising that such a diverse set of all-α protein structures can be generated simply by combining typical or canonical components of naturally occurring proteins,” said Dr. Koya Sakuma, a former Ph.D. student in SOKENDAI (The Graduate University for Advanced Studies).
The team selected five unique all-α proteins, each with five or six α-helices and distinguished by their complicated spatial arrangements, from the computationally generated structures. They then carried out de novo design of amino acid sequences that are capable of folding into the selected five all-α protein structures.
The experimental results were remarkable (See the figure). “The structures that emerged from structure determination process exhibit the complicated shapes, which perfectly matched computationally designed structures,” said Dr. Naohiro Kobayashi, a senior research fellow at RIKEN.
With this developed method, it is now possible to create all-α proteins with complicated shapes. Functions of proteins inherently depend on their three-dimensional structures. If we can design proteins with complicated shapes, it increases the potential to construct functional sites within them. This breakthrough research will pave the way for the creation of new functional proteins, which will contribute to healthcare and life sciences.
The research team includes Koya Sakuma from SOKENDAI (currently, Nagoya University), Naohiro Kobayashi from RIKEN, Toshihiko Sugiki from the Institute for Protein Research (IPR), Osaka University (currently Kitasato University), Toshio Nagashima from RIKEN, Toshimichi Fujiwara from IPR, Osaka University, Kano Suzuki from Chiba University, Naoya Kobayashi from ExCELLS NINS (currently, Nara Institute of Science and Technology), Takeshi Murata from Chiba University, Takahiro Kosugi from Institute for Molecular Science (IMS) NINS, Rie Tatsumi-Koga from ExCELLS NINS (currently IPR, Osaka University), Nobuyasu Koga from IMS NINS, ExCELLS NINS, SOKENDAI (currently IPR, Osaka University).
###
The article, “Design of complicated all-α protein structures”, was published in Nature Structural and Molecular Biologyat DOI: 10.1038/s41594-023-01147-9