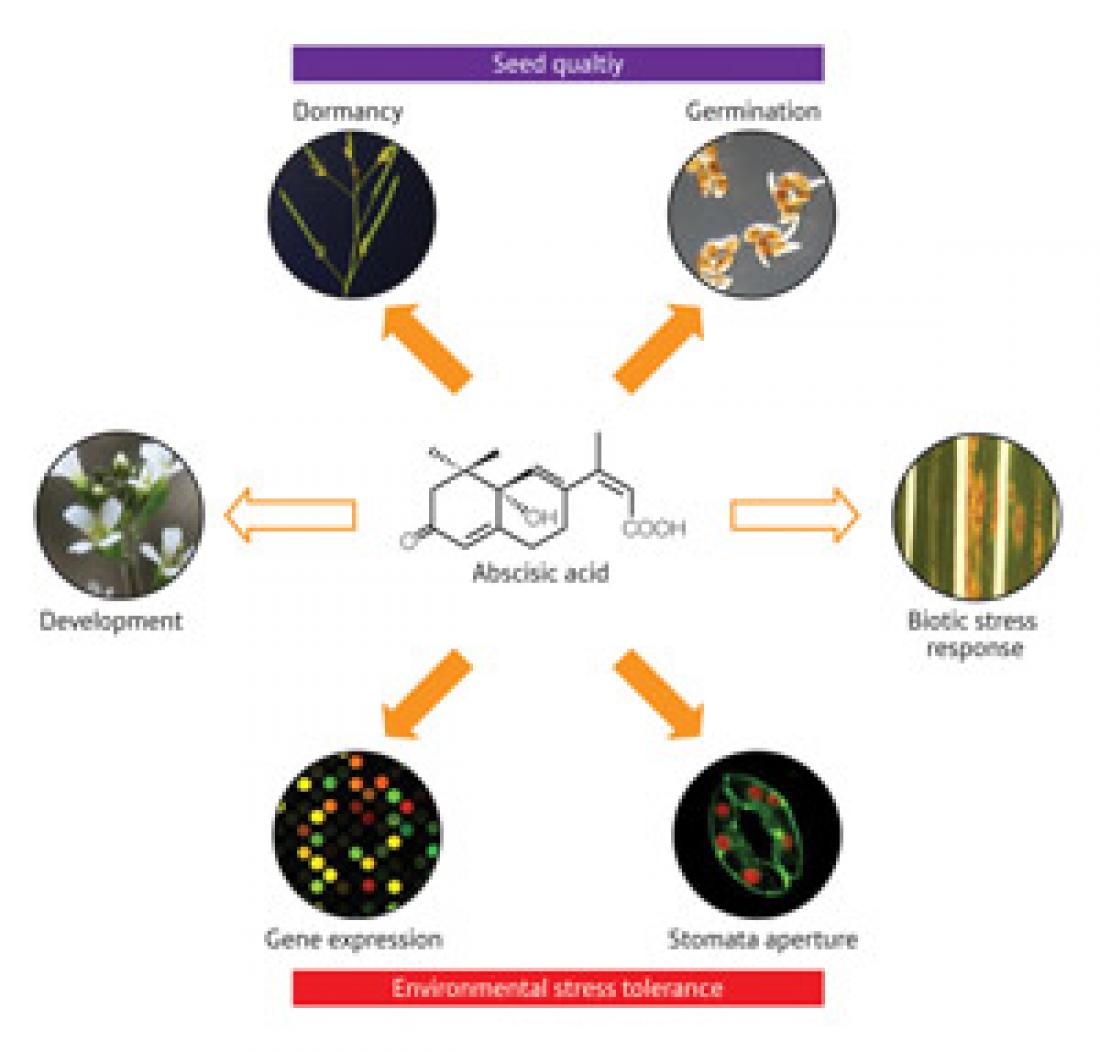
Schematic outlining the importance of the plant hormone abscisic acid (ABA), which controls a broad variety of crucial activities relating to plant development and survival.
Abcisic acid (ABA) has an awful lot on its plate. This plant hormone handles a staggeringly diverse array of jobs, from controlling seed germination to combating pathogen infection to dealing with drought conditions and scientists have assumed that the mechanisms regulating its activity must be equally elaborate. “ABA signal transduction pathways were thought to be a web-like, complex network in which many components have ‘some role’,” says Kazuo Shinozaki of the RIKEN Plant Science Center in Yokohama.
Taishi Umezawa of Shinozaki's team recently revealed the involvement of a subset of SNF1-related protein kinase 2 (SnRK2) proteins, which get activated via addition of phosphate groups in the aftermath of ABA signaling, enabling them to turn on downstream transcriptional activators. However, these signals also need to be turned off, and Takashi Hirayama’s team at the RIKEN Advanced Science Institute in Wako identified several protein phosphatase 2C (PP2C) enzymes that may constrain ABA signaling.
By combining their expertise, these two teams have now successfully sketched out the surprisingly simple processes underlying ABA regulation1. Thale cress plants lacking the three members (kinases) of SnRK2 subclass III are minimally responsive even to high ABA levels, highlighting their central role in this pathway. Subsequent experiments revealed that these kinases physically interact with a subset of PP2C enzymes, which directly remove the phosphate group; mutations that inactivate these PP2Cs lead in turn to SnRK2 hyperactivation.
Under normal conditions, PP2C appears to keep SnRK2 in a constant state of inactivation—but when ABA binds its receptor, the resulting complex interacts with PP2C in a manner that lifts this inhibition. Accordingly, the researchers found that mutant forms of PP2C that lack an ABA receptor-binding domain establish resistance to ABA signaling.
The investigators were taken aback by the simplicity of the model they identified. “We think this is the major ABA signaling pathway—and may even be the only ABA signaling pathway—[and it] consists of only four components: soluble ABA receptors, PP2C, SnRK2, and [downstream] transcription factors or other enzymes,” says Shinozaki. “That is very simple compared with our previous understanding.”
Of course, some additional levels of complexity still remain to be uncovered—most notably, understanding ABA’s part in the greater ecosystem of plant hormonal regulation. “We are interested in the molecular basis of cross-talk among plant hormone responses,” says Shinozaki. “Now that one major ABA signaling pathway is established, we can investigate this cross-talk at the molecular level.”
The corresponding author for this highlight is based at the Molecular Membrane Biology Laboratory, RIKEN Advanced Science Institute