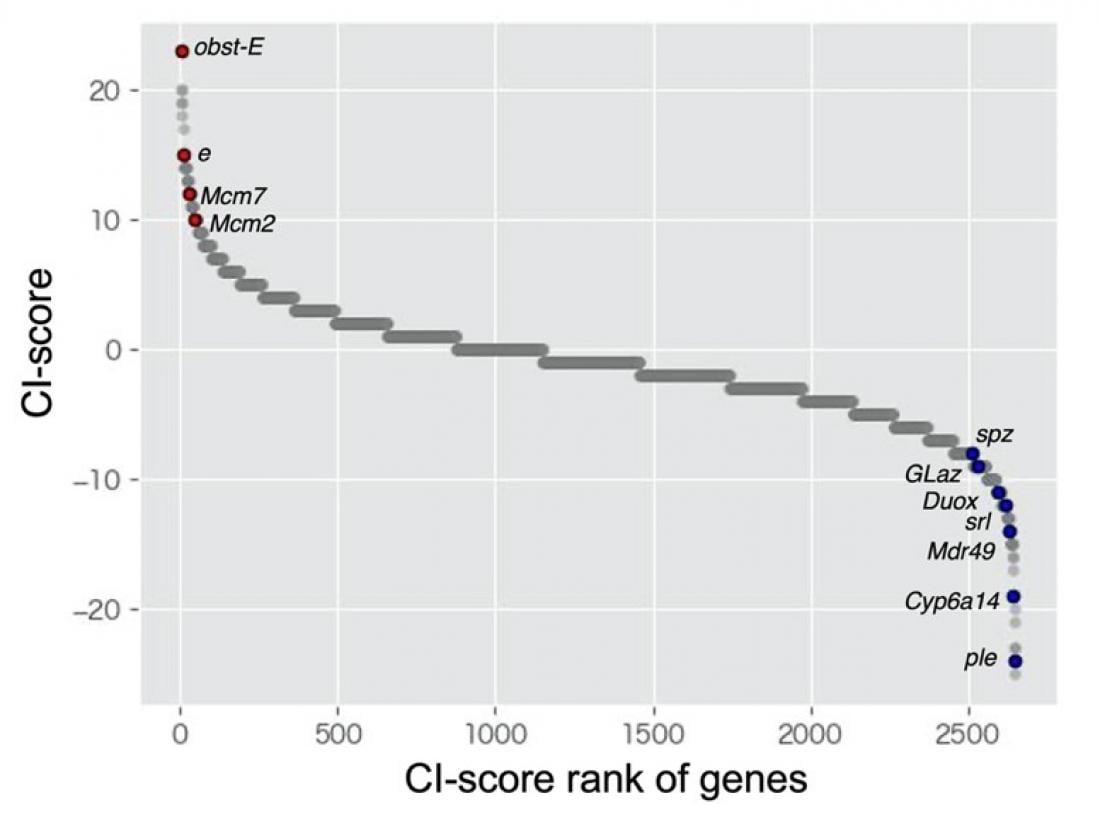
The horizontal axis indicates gene ranking based on the method developed in our laboratory for the meta-analysis of RNA-sequencing data. The vertical axis shows the values used to rank genes. High-ranking and low-ranking genes indicate upregulated genes in response to high and low densities, respectively. Gene names are described next to plots for previously reported density-response genes. In addition, the genes that have not been reported as density-responsive genes were also included in high-ranking and low-ranking genes.
A grasshopper hatched in a crowded environment may look and behave differently than a grasshopper hatched in isolation — even if they have the same genes. The mechanism of this density-dependent phenomenon, called polyphenism, is well-documented in both aphids and locusts, but how genes regulate these traits has remained shrouded until now. Researchers from Hiroshima University analyzed datasets collected from prior studies to better understand how genes can influence one another to change their expression depending on environmental conditions.
They published their results on Sept. 23 in Insects.
“Aphids exhibit multiple wing types and locusts exhibit different body colors and behaviors,” said corresponding author Hidemasa Bono, professor in Hiroshima University’s Graduate School of Integrated Sciences for Life. “These well-known agricultural pests are representative of insects that exhibit density-dependent plasticity. To reveal the molecules common to all or multiple species that exhibit this same type of plasticity, we collected and reanalyzed publicly available RNA sequencing data of aphids and locusts.”
The RNA sequencing data, called a transcriptome, is a collection of various expressed genes. It can also help identify new genes involved in producing specific traits. By performing a meta-analysis, researchers combine transcriptome results from multiple studies to see what the data says. In this study, the researchers analyzed 66 public transcriptome datasets from seven species of aphids and locusts.
“Meta-analysis is thought to be effective in providing additional insights into density-dependent polyphenism because it can uncover new information that would not be found with conventional hypothesis-driven research methods,” said first author Kouhei Toga, researcher in Hiroshima University’s Graduate School of Integrated Sciences for Life. “This study is the first meta-analysis conducted on datasets of two evolutionarily distant lineages, and it identified many density-responsive genes, which have scarcely been the focuses of research that aims to elucidate the molecular mechanisms of density-dependent plasticity.”
Specifically, the researchers found that DNA replication, DNA metabolic processes and the mitotic cell cycle were all enriched in response to crowded conditions. According to Toga, their results emphasize the importance of these processes— which have rarely been the focus of research in this area — as regulatory mechanisms in density-dependent polyphenism research.
They also found discrepancies with some studies, including one that found a gene related to pigmentation in more gregarious locusts was more highly expressed in isolated conditions. When compared with the data from other studies, the researchers found the gene fit in a category of other genes that upregulated their expression when under oxidative stress. According to Bono, oxidative stress is a more likely explanation for the high gene expression in solitary locusts than in crowded conditions.
“We also found neurological system modifications may play an important role in inducing density-dependent phenotypic changes in two lineages,” Bono said, explaining that several genes functioning in the nervous system, which would lead to density-dependent behavioral changes, increased under isolated conditions.
The findings can be generally applied to other species that exhibit density-dependent polyphenism due to the sheer amount of data from so many studies, which serves as a cross-check of previous hypotheses and results, according to Toga.
“With increasing public RNA sequencing data, a meta-analysis that combines the data from multiple studies has succeeded in providing novel insights into targeted biological processes,” Toga said. “We hope that functional analysis of the genes identified in this study will lead to the development of methods to control the growth of aphids and locusts. We also hope to clarify how organisms respond and adapt to density by applying meta-analysis to various species.”
###
The Center for Innovation for Bio-Digital Transformation and the Japan Science and Technology Agency supported this research. Computations for this work were performed on the computers at the Hiroshima University Genome Editing Innovation Center.
About Hiroshima University
Since its foundation in 1949, Hiroshima University has striven to become one of the most prominent and comprehensive universities in Japan for the promotion and development of scholarship and education. Consisting of 12 schools for undergraduate level and 4 graduate schools, ranging from natural sciences to humanities and social sciences, the university has grown into one of the most distinguished comprehensive research universities in Japan.
English website: https://www.hiroshima-u.ac.jp/en