To facilitate the interpretation of experimental imaging, Romain Amyot and Holger Flechsig from NanoLSI have now developed the mathematical framework and computational methods to reconstruct 3D atomistic structures from AFM surface scans. In their recent research paper [1], which is a collaboration with experimental researchers, they provide applications to high-speed AFM images ranging from single molecular machines, protein filaments, to even large-scale assemblies of protein lattices, and demonstrate how the full atomistic information advances the molecular understanding beyond topographic images.
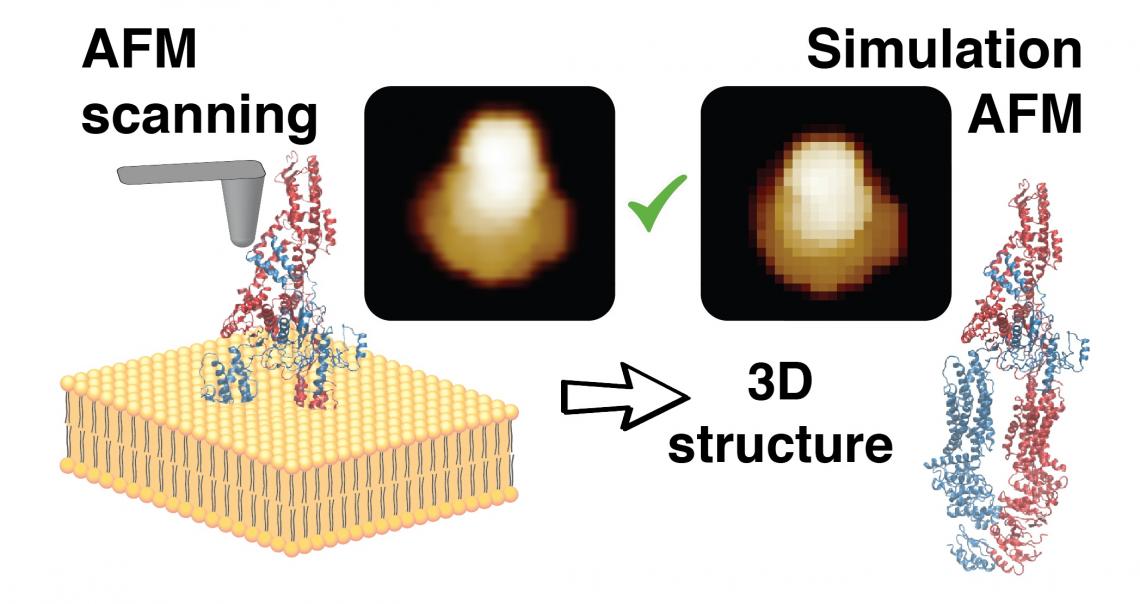
The developed methods employ simulation AFM, which was previously established by Amyot and Flechsig, and distributed within the free BioAFMviewer software package (PLoS Comput Biol, 2020). Simulation AFM computationally emulates experimental scanning of biomolecules to translate structural data into simulation AFM topographic images that can be compared to real AFM images. A procedure of automatized fitting was then implemented to identify the high-resolution molecular structure behind a limited-resolution experimental AFM image. It is therefore possible to retrieve full 3D atomistic information from just a scan of the protein surface obtained under AFM observations. To illustrate the explanatory potential of this achievement, Flechsig describes, “Imagine that instead of just seeing the tip of an iceberg, you are now able to see everything hidden under the sea, to the extent that you can even detect impurities or density differences within its structure, helping you to explain the icebergs’ coloration.”
To share the developments with the broad Bio-AFM community, all computational methods are embedded within the user-friendly BioAFMviewer interactive software interface. The new methods are already applied in numerous interdisciplinary collaborations to help understand experimental observations.
Illustration of the method to reconstruct the 3D atomistic protein structure from an experimental AFM surface scan.
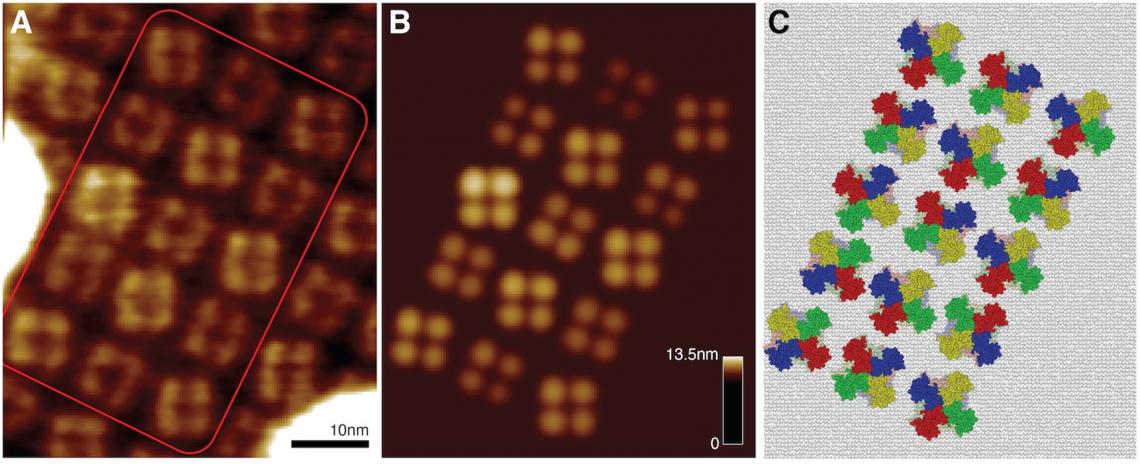
From the membrane channel lattice HS-AFM image (left), the optimal-match simulation-AFM lattice (middle) provides the atomic-resolution arrangement of proteins (right). This information allows a molecular-level explanation of how these protein channels become activated.