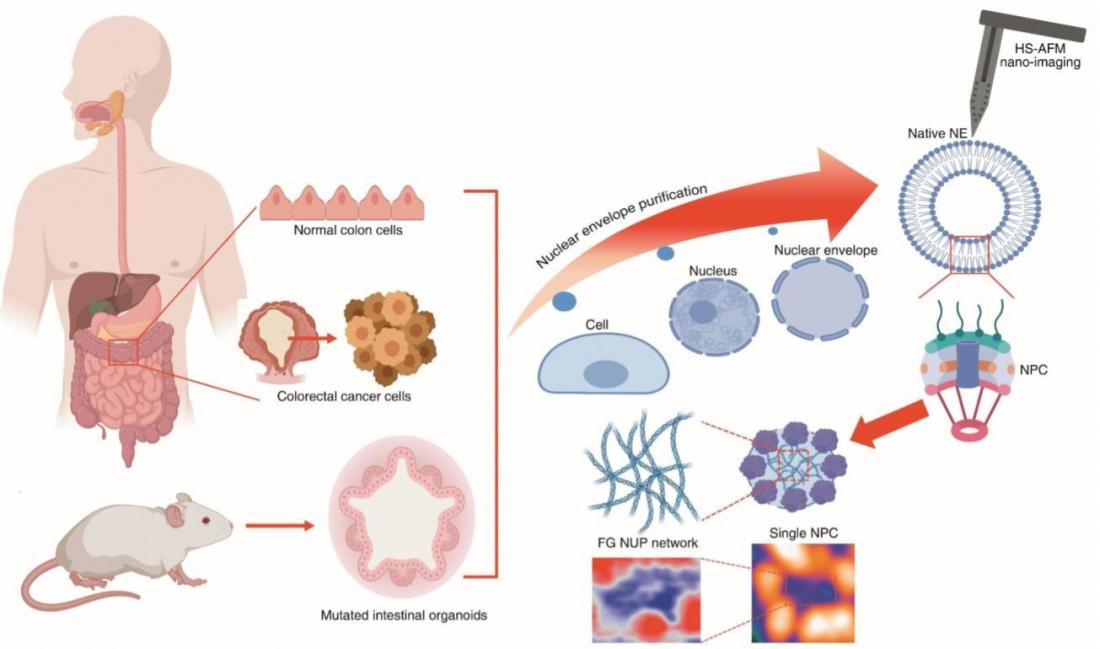
Figure1. Schematic illustration of manipulation and tracking of native nuclear nano-pores from colon cancer cells and mouse organoids. Live tracking of single bio-filament conformations inside the nuclear pore presented as a nano diagnosis tool for the colorectal cancer.
In human cells, the nucleus is enclosed by a structure called the nuclear pore complex (NPC). It acts as a ‘gatekeeper’ controlling the transport of molecules between the nucleus and the surrounding cytoplasm (the protein-containing solution in the inside of a cell). The NPC consists of proteins known as nucleoporins; some of these, the so-called FG-NUPs, belong to the class of intrinsically disordered proteins (IDPs) and capable of forming liquid-‐liquid phase separation (LLPS), lacking a well-defined tertiary structure (that is, a particular 3D shape). Although a lot is known about FG-NUPs, a thorough understanding of how their structure varies in time and space has been missing. But now, by applying high-speed atomic force microscopy (HS-AFM), Richard Wong from Kanazawa University and colleagues provide much-needed insights into the spatiotemporal structure of FG-NUPs.
The technique used by the researchers, HS-AFM, is typically used for imaging surfaces. A tiny cantilever is made to move over the surface; at any given time, the force experienced by the cantilever probe can be converted into a height measure. A scan of the whole surface then results in a height map of the sample. By repeatedly scanning the surface rapidly, a video of its evolving structure is obtained. Applying HS-AFM to FG-NUPs, Wong and colleagues were able to measure several of the molecules’ properties, including the extension velocity of FG-NUP filaments (thread-like protruding structures), their bending angles and how they form knots.
The scientists studied FG-NUPs in normal colon cells and in colorectal cancer cells and organoids. They found that the former displayed less conformational dynamics. A particularly interesting conclusion is that in colon cancer cells, the structure of the so-called central plug is smaller, and cannot develop filamentous features as easily as in normal cells, a finding with high clinical relevance.
The results of Wong and colleagues regarding the central plug are very important and timely, as its morphology and function have been the subject of debate. The researchers now provide strong evidence that the central plug at least partially consists of FG-NUPs.
Apart from demonstrating that HS-AFM is a tool capable of visualizing FG-NUP filament motion in real time, another implication of the work of the scientists is “that bio-recycled nanomaterials [like NPC nanopores] … have biocompatible advantages … directly derived from cells and organoids, rather than other engineered nanomaterials [like e.g. carbon nanotubes, which may induce tumors and related pathologies] opening a new avenue for nano-tissue engineering.”
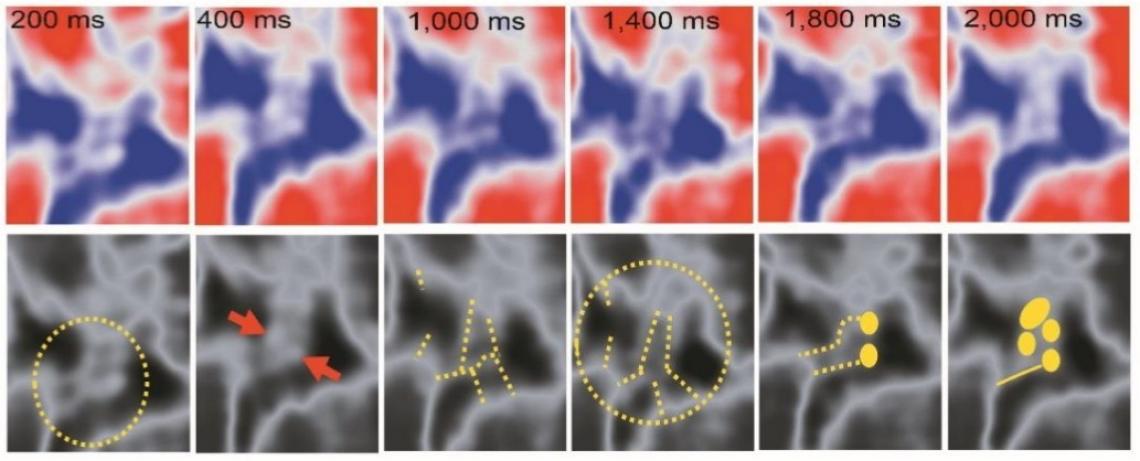
Figure2. HS-AFM enables dynamically studying filament structures in FG-NUP LLPS proteins, including twisting and knot formation.
[Background]
Nuclear pore complex
The nucleus of a cell is of key importance to any organism. It stores and organizes genetic information (DNA) in a way separating it from other cellular components in the surrounding cytoplasm. The nuclear pore complex (NPC), a very large protein complex dressed around the nucleus, is the ‘gatekeeper’ in the exchange of molecules between the nucleus and the cytoplasm; it lets material pass that should reach the nucleus and blocks material that should not. This communication can happen because of pores in the NPC, structures built from proteins known as FG-NUPs. FG-NUPs do not have well-defined shapes; instead, they vary in time and space. By applying a technique called high-speed atomic force microscopy, Richard Wong from Kanazawa University and colleagues have now provided new, valuable insights into the spatiotemporal structure of FG-NUPs of both normal and cancer cells.
Atomic force microscopy
Atomic force microscopy (AFM) is an imaging technique in which the image is formed by scanning a surface with a very small tip. Horizontal scanning motion of the tip is controlled via piezoelectric elements, while vertical motion is converted into a height profile, resulting in a height distribution of the sample’s surface. As the technique does not involve lenses, its resolution is not restricted by the so-called diffraction limit. In a high-speed setup (HS-AFM), the method can be used to produce movies of a sample’s structural evolution in real time. Wong and colleagues have successfully used HS-AFM to study the dynamics of FG-NUPs, proteins playing a key role in the transport-regulating function of the nuclear pore complex situated between a cell’s nucleus and the surrounding cytoplasm.