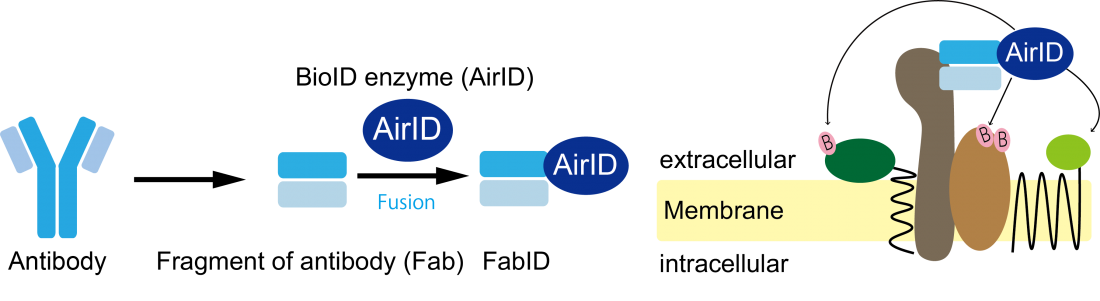
Innovative technology for the comprehensive analysis of membrane protein extracellular interactions.
1. Background
Many proteins within the body form complexes with other proteins, determining the fate of cells. Therefore, the analysis of protein-protein interactions (PPI) is a crucial process for understanding the biological function of target proteins. Membrane proteins, which account for over 30% of human genes, play a vital role in cell functions. Many membrane proteins are known to form complexes to exert their functions, making the elucidation of membrane protein PPIs essential for understanding protein function. However, the development of technologies to analyze PPIs, especially extracellular protein–protein interactions (exPPIs) of living cells, has been lagging. Recently, proximity labeling methods that label proteins in close proximity for large-scale PPI analysis have gained attention. However, proximity protein labeling methods targeting the extracellular region of membrane proteins have primarily involved PPI analysis using molecules with cellular toxicity, prompting a search for systems targeting living cells.
2. Research Findings
The Proteo-Science Center at Ehime University has independently developed the proximity-dependent biotin labeling enzyme AirID (Kido, et al., eLife 2020), which biotinylates lysine residues of proteins in close proximity. There have been several studies using proximity-dependent biotin labeling enzymes for exPPI analysis, but they involved genetically modified proteins with significantly different shapes from their original structures, making it unclear to what extent the analysis results reflect the original interactions. To accurately understand interactions occurring on the cell membrane of living cells, it was necessary to develop a technology that could directly target proteins expressed by the cells for exPPI analysis. Therefore, our research group initiated this study on the premise that it would be possible to analyze exPPI by creating a molecule (FabID) that fuses AirID to the antigen recognition site of an antibody recognizing the extracellular domain of membrane proteins.
The membrane protein analyzed for exPPI was the epidermal growth factor receptor EGFR, a protein known as a cancer gene located on the cell membrane. When FabID and biotin were added to epithelial-like carcinoma-derived cells (A431 cells) (hereinafter referred to as epithelial cancer cells) expressing EGFR, it was confirmed at the cell culture level that EGFR on the cell membrane could be biotinylated using FabID.
By combining biotin labeling with FabID and mass spectrometry developed at Tokushima University for analyzing biotinylated proteins, we successfully identified many novel EGFR interacting proteins. The identified proteins have the potential to be new drug targets.
EGFR transmits signals to cells by binding to a ligand known as EGF. It is known that when EGF binds to EGFR, various proteins bind to the intracellular domain of EGFR, forming a protein complex. Currently, EGFR tyrosine kinase inhibitors, widely used as cancer treatments, are thought to exert their therapeutic effects by binding to EGFR and inhibiting the formation of protein complexes mediated by the EGFR intracellular domain. However, the ligand-dependent and drug-dependent exPPI changes of EGFR when EGFR binds to EGF (ligand) or when EGFR tyrosine kinase inhibitors (drugs) act have not been observed at all. Therefore, we observed ligand-dependent and drug-dependent exPPI changes of EGFR occurring in cultured cells using FabID. As a result, we are the first in the world to find that exPPI changes dynamically when ligands and drugs bind to EGFR.
3. Ripple Effect
Membrane proteins are used by almost all organisms to transmit information inside and outside cells. Therefore, analyzing the exPPI of membrane proteins is expected to directly contribute to the advancement of receptor biology. Moreover, since a majority of commercial drugs function by targeting membrane proteins, membrane proteins are known as important drug targets. However, finding new membrane protein drug targets is challenging, and it has been a significant issue in the pharmaceutical industry. The FabID technology developed in this study not only enables exPPI analysis using living cells but can also be used to identify novel drug targets. Furthermore, FabID has been found to capture ligand-dependent and drug-dependent exPPI changes that have not been analyzed by conventional methods. In the future, the identification of new drug targets through exPPI analysis using FabID and detailed analysis of membrane protein exPPI changes when commercial drugs bind are expected to contribute significantly to the development of commercial drugs.
4. Research Organization and Support
This research was conducted as a joint research project by the Proteo-Science Center at Ehime University, Institute of Advanced Medical Sciences at Tokushima University, the School of Life Sciences at Tokyo Pharmaceutical University, Department of Molecular Pharmacology of Tohoku University Graduate School of Medicine, and the Department of Bioscience of Nagahama Institute of BioScience and Technology.